To evaluate the relationship between antipseudomonal antibiotic consumption and each individual drug resistance rate in Pseudomonas aeruginosa strains causing ICU acquired invasive device-related infections (IDRI).
DesignA post hoc analysis was made of the data collected prospectively from the ENVIN-HELICS registry.
SettingIntensive Care Units participating in the ENVIN-UCI registry between the years 2007 and 2016 (3-month registry each year).
PatientsPatients admitted for over 24h.
Main variablesAnnual linear and nonlinear trends of resistance rates of P. aeruginosa strains identified in IDRI and days of treatment of each antipseudomonal antibiotic family per 1000 occupied ICU bed days (DOT) were calculated.
ResultsA total of 15,095 episodes of IDRI were diagnosed in 11,652 patients (6.2% out of a total of 187,100). Pseudomonas aeruginosa was identified in 2095 (13.6%) of 15,432 pathogens causing IDRI. Resistance increased significantly over the study period for piperacillin-tazobactam (P<0.001), imipenem (P=0.016), meropenem (P=0.004), ceftazidime (P=0.005) and cefepime (P=0.015), while variations in resistance rates for amikacin, ciprofloxacin, levofloxacin and colistin proved nonsignificant. A significant DOT decrease was observed for aminoglycosides (P<0.001), cephalosporins (P<0.001), quinolones (P<0.001) and carbapenems (P<0.001).
ConclusionsNo significant association was observed between consumption of each antipseudomonal antibiotic family and the respective resistance rates for P. aeruginosa strains identified in IDRI.
Evaluar la relación entre el consumo de antibióticos antipseudomonales y la tasa de resistencia de cada fármaco individual en cepas de Pseudomonas aeruginosa aisladas en infecciones relacionadas con dispositivos invasivos (IDRI, por sus siglas en inglés) adquiridas en la unidad de cuidados intensivos (UCI).
DiseñoAnálisis post-hoc de los datos recopilados prospectivamente del registro ENVIN-HELICS.
ÁmbitoLas UCI que participaron en el registro ENVIN-UCI entre los años 2007-2016 (registro de 3 meses cada año).
PacientesPacientes ingresados >24h.
Variables principalesSe calcularon las tendencias anuales lineales y no lineales de las tasas de resistencia de las cepas de P. aeruginosa identificadas en IDRI y los días de tratamiento de cada familia de antibióticos antipseudomonales por 1.000 días de cama ocupada en la UCI (DOT).
ResultadosSe diagnosticaron 15.095 episodios de IDRI en 11.652 pacientes (6,2% de 187.100). Se identificó P. aeruginosa en 2.095 (13,6%) de 15.432 patógenos que causaron IDRI. La resistencia aumentó significativamente durante el período de estudio para piperacilina-tazobactam (p<0,001), imipenem (p=0,016), meropenem (p=0,004), ceftazidima (p=0,005) y cefepima (p=0,015), mientras que las variaciones en las tasas de resistencia de amikacina, ciprofloxacina, levofloxacina y colistina no fueron significativas. Se observó una disminución significativa de la DOT para aminoglucósidos (p<0,001), cefalosporinas (p<0,001), quinolonas (p<0,001) y carbapenems (p<0,001).
ConclusionesNo se encontró asociación significativa del consumo de cada familia de antibióticos antipseudomonales con sus respectivas tasas de resistencia para las cepas de P. aeruginosa identificadas en IDRI.
The relationship between antimicrobial use and increasing emergence of resistance to specific agents or antimicrobial families has been well documented.1–11 In the ICU setting, other factors unrelated to pressure of antimicrobial use may account for the alarming rate of colonized or infected patients by multiresistant microoganisms (MRB), such as admission of patients already colonized or infected,12 spread by cross-transmission from MRB carriers,13 and especially contaminated hospital reservoirs.14–17
Pseudomonas aeruginosa is frequently implicated in invasive device-related infection (IDRI) especially in ICUs.18,19 The increasing resistance to antipseudomonal antimicrobials and the occurrence of epidemic outbreaks of MDR P. aeruginosa has become a challenging clinical problem in ICU populations.12,17
In Spain, data of IDRI diagnosed in ICU patient are collected since 1994 in the National ICU-Acquired Infection Surveillance Study (ENVIN-HELICS registry) database. Annual reports of the ENVIN-HELICS surveillance program provide detailed information on etiology of IDRI and antimicrobial consumption. The aim of this study was evaluate the relationship between antipseudomonal antibiotic consumption and each individual drug's resistance rate in P. aeruginosa strains causing ICU invasive device-related infections (IDRI). It was hypothesized that there was a relationship between ICU consumption of antipseudomonal antimicrobials and development of P. aeruginosa resistant strains.
MethodsDesign and study populationThis was a retrospective analysis of data collected prospectively from the ENVIN-HELICS registry in the framework of an observational, nationwide, multicenter study. The purpose of ENVIN-HELICS database is to register the frequency, etiology, and presence of MRB in ICU patients with IDRI. Invasive device-related infections include ventilator-associated pneumonia (VAP), catheter-associated urinary tract infection (CAUTI) and primary bacteremia including catheter-related bloodstream infection (CRBSI) and bacteremia of unknown origin. Indications and days of treatment of all antimicrobials used during the patients’ ICU stay are also registered. All patients admitted for more than 24h to the participating ICUs during a 3-month period (between April 1st and June 30th) over 10 consecutive years (2007–2016) were included in the study provided that the diagnosis of IDRI related to invasive devices caused by P. aeruginosa had been established during the patient's stay in the ICU.
The ENVIN-HELICS registry was developed by the Study Group of Infectious Diseases and Sepsis (GTEIS) of the Spanish Society of Intensive Care Medicine and Coronary Units (SEMICYUC) in 1994. Data from about 200 ICUS (about 80% of the total ICUs in Spain) are collected using the ENVIN-HELICS software application located in a web-based server available at http://hws.vhebron.net/ENVIN-helics.19 Participation in the registry is voluntary and data collection is longitudinal and prospective. Quality-control audits ensured internal quality of the clinical information recorded in the database.20
The ENVIN registry was approved by the Ethics Committees of the participating ICUs and was declared a registry of healthcare interest by the Spanish Ministry of Health, Social Services and Equality in 2014. A consent statement was not applicable due to the non-interventional nature of the study because data were collected from the ENVIN-HELICS registry.
Study variablesStudy variables included the annual rate of P. aeruginosa resistance to antipseudomonal antibiotics used in ICU patients and days of treatment per 1000 occupied bed-days (DOT), which were calculated for each antipseudomonal agent and antimicrobial families annually as well as in the same year and the previous year for all ICUs that participated in the registry. All P. aeruginosa isolates from VAP, CAUTI, and primary bacteremia (bacteremia of unknown origin and/or [CRBSI]) in patients with an established central venous catheter were recorded. Definitions of these infections were those reported in the manual of the ENVIN project following indications published by the European Centre for Disease Control and Prevention.21,22 Infections associated with invasive devices were diagnosed by attending physicians and recorded in the patient's medical history. Physicians responsible for surveillance of nosocomial infections were intensivists with special interest in infectious diseases.
Susceptibility of P. aeruginosa isolates to different antimicrobials was assessed at the Services of Clinical Microbiology of the participating hospitals, in many of them following specifications (method and breakpoints) of the European Committee on Antimicrobial Susceptibility Testing.23 Antipseudomonal categories and antimicrobials included in each category were as follows: aminoglycosides (amikacin), carbapenems (imipenem and meropenem), cephalosporins (ceftazidime and cefepime), quinolones (levofloxacin and ciprofloxacin), ureidopenicillins (piperacillin-tazobactam), and colistin.
Frequency measuresThe resistance rate to each antipseudomonal agent was expressed in percentages and calculated as the number of resistant strains divided by the total number of P. aeruginosa strains identified in each annual period for which antimicrobial susceptibility testing against this agent was available, per 100. DOTs were determined for individual agents and for all antimicrobials of each antipseudomonal families (aminoglycosides, carbapenems, cephalosporins, quinolones, and ureidopenicillins).
Statistical analysisStatistical analyses included estimated annual linear and non-linear trends of resistance rates and DOT per antimicrobial (period 2007 to 2016). Resistance and DOT relationship was adjusted by family of antipseudomonal antimicrobials of the current year (period 2007 to 2016) and by the family DOT of the current and the previous year (period 2008 to 2016). A Poisson regression model was used for adjustment and the 95% confidence intervals (CIs) were calculated. Statistical significance was set at P<0.05. Data were analyzed with the R program.24
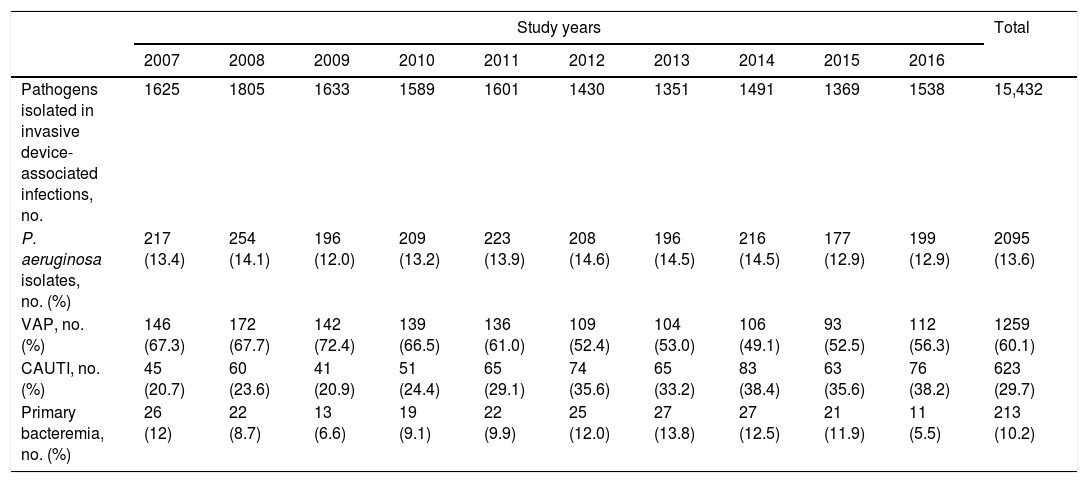
ResultsDuring the study period, a total 187,100 patients with 1,440,472 patient-days of ICU stay (mean 7.7 days) were included in the ENVIN-HELICS registry. A total of 15,095 episodes of IDRI were diagnosed in 11,652 patients (6.2% of a total of 187,100). P. aeruginosa was identified in 2095 (13.6%) of 15,432 pathogens causing IDRI. Table 1 shows the overall number of IDRI caused by P. aeruginosa for each study year, as well as the corresponding numbers of VAP, CAUTI, and primary bacteremia. P. aeruginosa mostly caused VAP, although with a trend to decrease in recent years (from 67.3% in 2007 to 56.3% in 2016), whereas an increase of CAUTI caused by P. aeruginosa was observed (from 20.7% in 2007 to 38.2% in 2016).
Evolution of Pseudomonas aeruginosa strains identified each year in HCAIs related to invasive devices controlled in the ENVIN-HELICS registry.
| Study years | Total | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 2007 | 2008 | 2009 | 2010 | 2011 | 2012 | 2013 | 2014 | 2015 | 2016 | ||
| Pathogens isolated in invasive device-associated infections, no. | 1625 | 1805 | 1633 | 1589 | 1601 | 1430 | 1351 | 1491 | 1369 | 1538 | 15,432 |
| P. aeruginosa isolates, no. (%) | 217 (13.4) | 254 (14.1) | 196 (12.0) | 209 (13.2) | 223 (13.9) | 208 (14.6) | 196 (14.5) | 216 (14.5) | 177 (12.9) | 199 (12.9) | 2095 (13.6) |
| VAP, no. (%) | 146 (67.3) | 172 (67.7) | 142 (72.4) | 139 (66.5) | 136 (61.0) | 109 (52.4) | 104 (53.0) | 106 (49.1) | 93 (52.5) | 112 (56.3) | 1259 (60.1) |
| CAUTI, no. (%) | 45 (20.7) | 60 (23.6) | 41 (20.9) | 51 (24.4) | 65 (29.1) | 74 (35.6) | 65 (33.2) | 83 (38.4) | 63 (35.6) | 76 (38.2) | 623 (29.7) |
| Primary bacteremia, no. (%) | 26 (12) | 22 (8.7) | 13 (6.6) | 19 (9.1) | 22 (9.9) | 25 (12.0) | 27 (13.8) | 27 (12.5) | 21 (11.9) | 11 (5.5) | 213 (10.2) |
VAP: ventilator-associated pneumonia; CAUTI: catheter-associated urinary tract infection; no.: number.
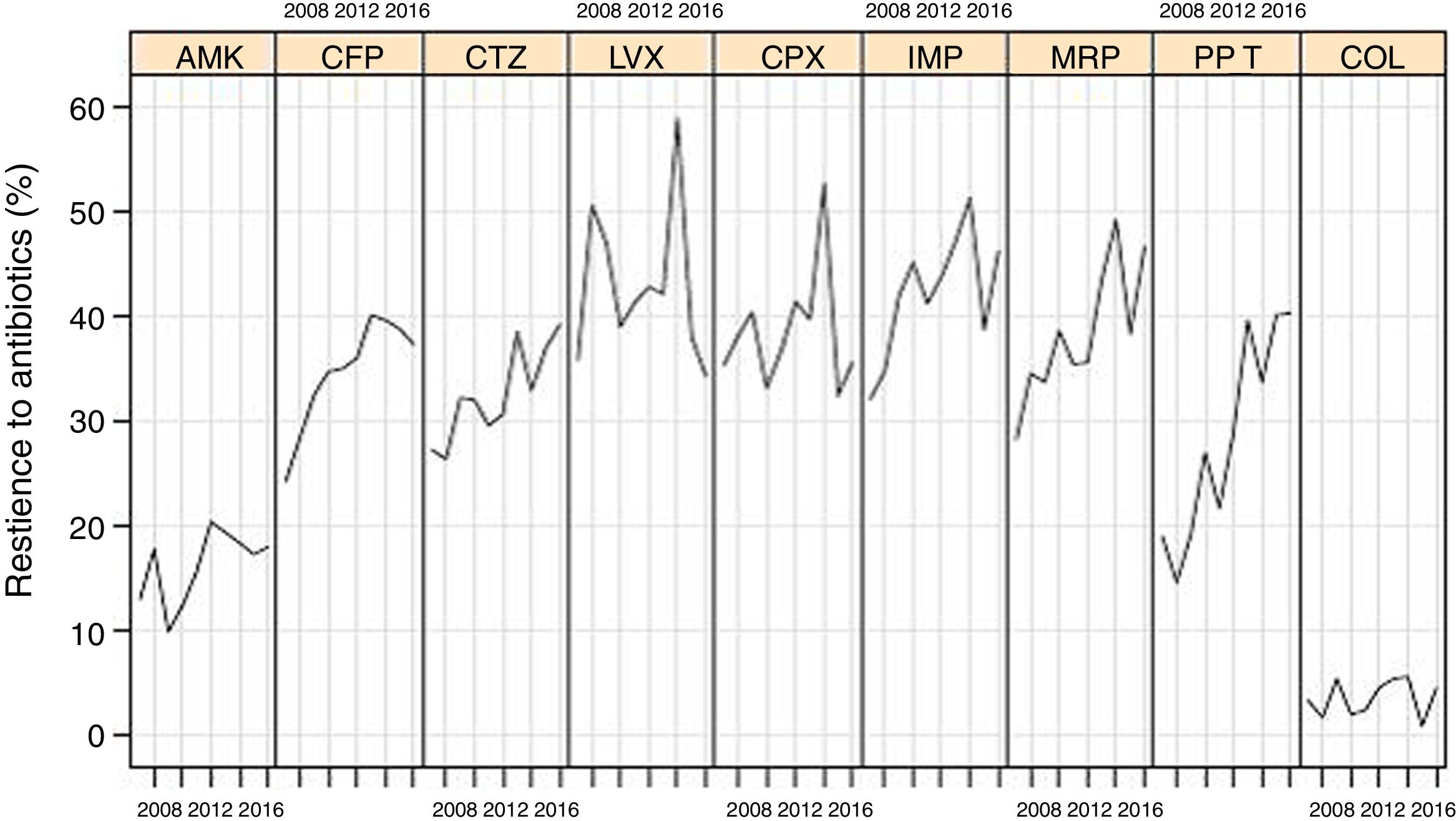
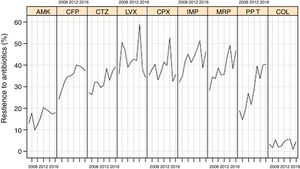
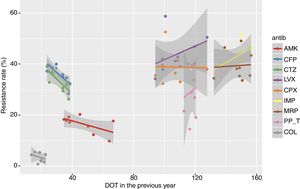
Resistance rates of P. aeruginosa strains to different antipseudomonal antimicrobials during the study period are shown in Fig. 1. There was a progressive increase of resistance to piperacillin-tazobactam (from 18.9% in 2007 to 40.2% in 2016, P<0.001), imipenem (from 32.0% to 46.1%, P=0.016), meropenem (from 28.2% to 46.5%, P=0.004), ceftazidime (from 27.2% to 39.1%, P=0.005), cefepime (from 24.2% to 37.2%, P=0.015), and to a lower extent for amikacin (from 12.9% to 17.9%, P=0.084). Resistance rates to ciprofloxacin (from 35.2% to 35.5%), levofloxacin (from 35.7% to 34.2%), and colistin (from 3.2% to 4.4%) remained stable.
Linear evolution of antipseudomonal antimicrobials resistance rates of P. aeruginosa isolated in invasive device-related infections in ICU patients between 2007 and 2016. (AMK: amikacin; CFP: cefepime; CTZ: ceftazidime; LVX: levofloxacin; CPX: ciprofloxacin; IMP: imipenem; MRP: meropenem; PP_T: piperacillin-tazobactam; COL: colistin).
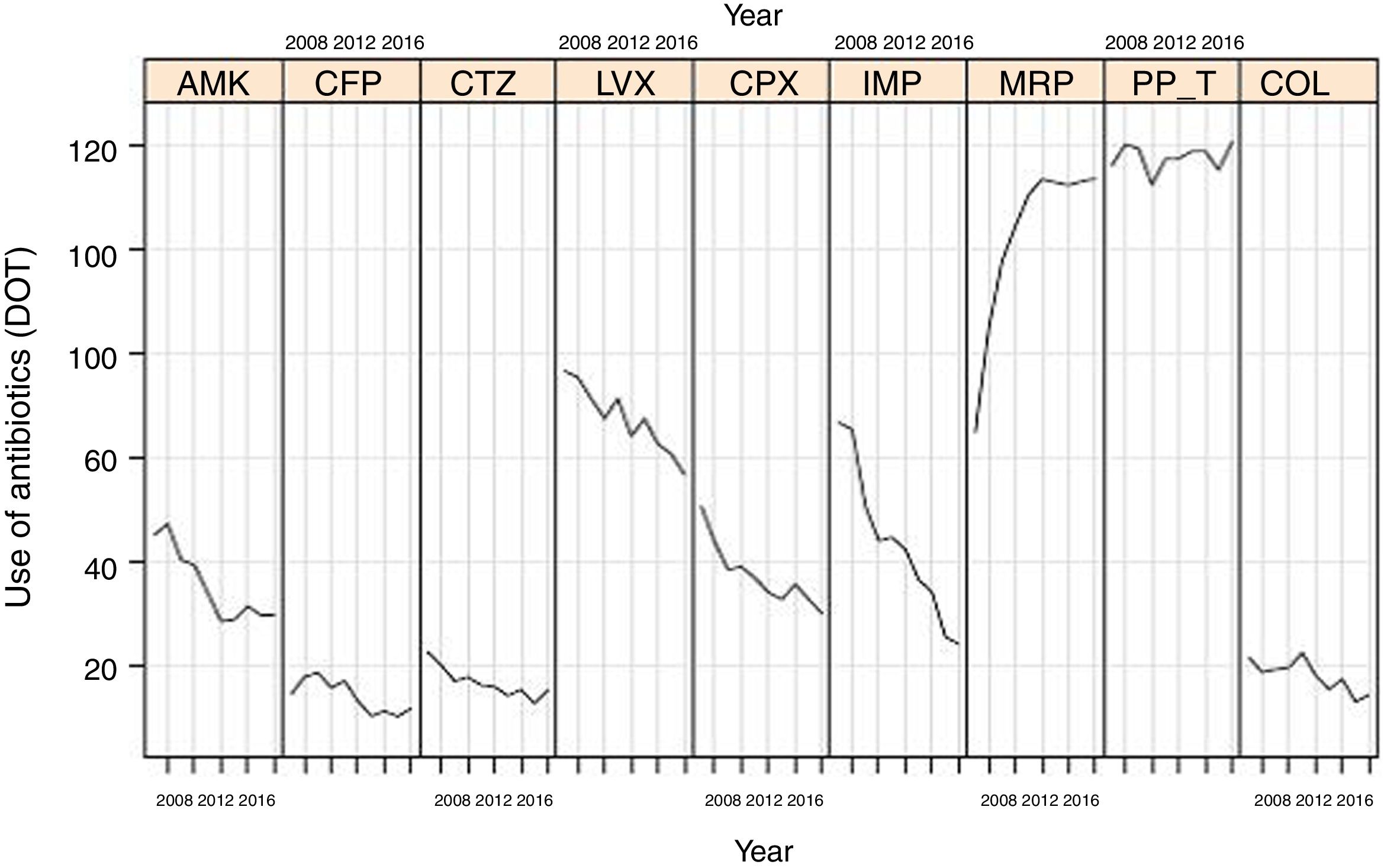
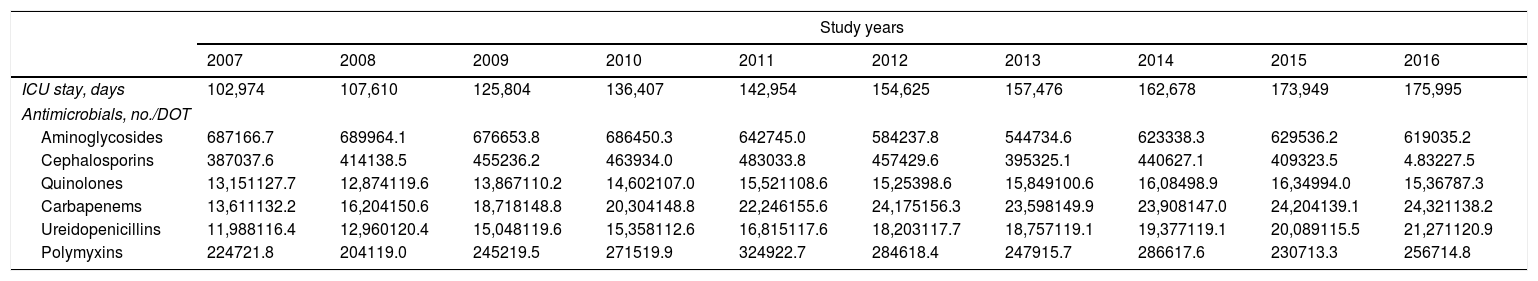
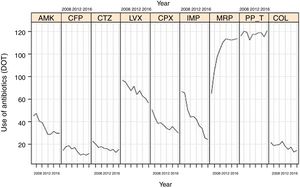
Evolution of consumption of individual antipseudomonal antimicrobials over 10 years is shown in Table 2. Between 2007 and 2016, a significant decrease of DOT for aminoglycosides (from 66.7 to 35.2, P<0.001), cephalosporins (from 37.6 to 27.5, P<0.001), quinolones (from 127.7 to 87.3, P<0.001), and carbapenems (from 150.6 to 138.2, P<0.001) was observed. The consumption of ureidopenicillins remained stable (from 116.4 in 2007 to 120.9 in 2016, P=0.096). Changes of DOT for each antipseudomonal antibiotic are shown in Fig. 2. There was an important predominance of beta-lactam antibiotic families, with common mechanisms of action and development of resistances.
Evolution of each antipseudomonal antimicrobial family consumption during the study period (2007–2016) expressed as DOT (days of treatment/days of ICU stay×1000).
| Study years | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 2007 | 2008 | 2009 | 2010 | 2011 | 2012 | 2013 | 2014 | 2015 | 2016 | |
| ICU stay, days | 102,974 | 107,610 | 125,804 | 136,407 | 142,954 | 154,625 | 157,476 | 162,678 | 173,949 | 175,995 |
| Antimicrobials, no./DOT | ||||||||||
| Aminoglycosides | 687166.7 | 689964.1 | 676653.8 | 686450.3 | 642745.0 | 584237.8 | 544734.6 | 623338.3 | 629536.2 | 619035.2 |
| Cephalosporins | 387037.6 | 414138.5 | 455236.2 | 463934.0 | 483033.8 | 457429.6 | 395325.1 | 440627.1 | 409323.5 | 4.83227.5 |
| Quinolones | 13,151127.7 | 12,874119.6 | 13,867110.2 | 14,602107.0 | 15,521108.6 | 15,25398.6 | 15,849100.6 | 16,08498.9 | 16,34994.0 | 15,36787.3 |
| Carbapenems | 13,611132.2 | 16,204150.6 | 18,718148.8 | 20,304148.8 | 22,246155.6 | 24,175156.3 | 23,598149.9 | 23,908147.0 | 24,204139.1 | 24,321138.2 |
| Ureidopenicillins | 11,988116.4 | 12,960120.4 | 15,048119.6 | 15,358112.6 | 16,815117.6 | 18,203117.7 | 18,757119.1 | 19,377119.1 | 20,089115.5 | 21,271120.9 |
| Polymyxins | 224721.8 | 204119.0 | 245219.5 | 271519.9 | 324922.7 | 284618.4 | 247915.7 | 286617.6 | 230713.3 | 256714.8 |
Linear evolution of consumption of antipseudomonal antimicrobials (expressed as DOT) in Spanish ICUs participating in the ENVIN-HELICS registry. (AMK: amikacin; CFP: cefepime; CTZ: ceftazidime; LVX: levofloxacin; CPX: ciprofloxacin; IMP: imipenem; MRP: meropenem; PP_T: piperacillin-tazobactam; COL: colistin).
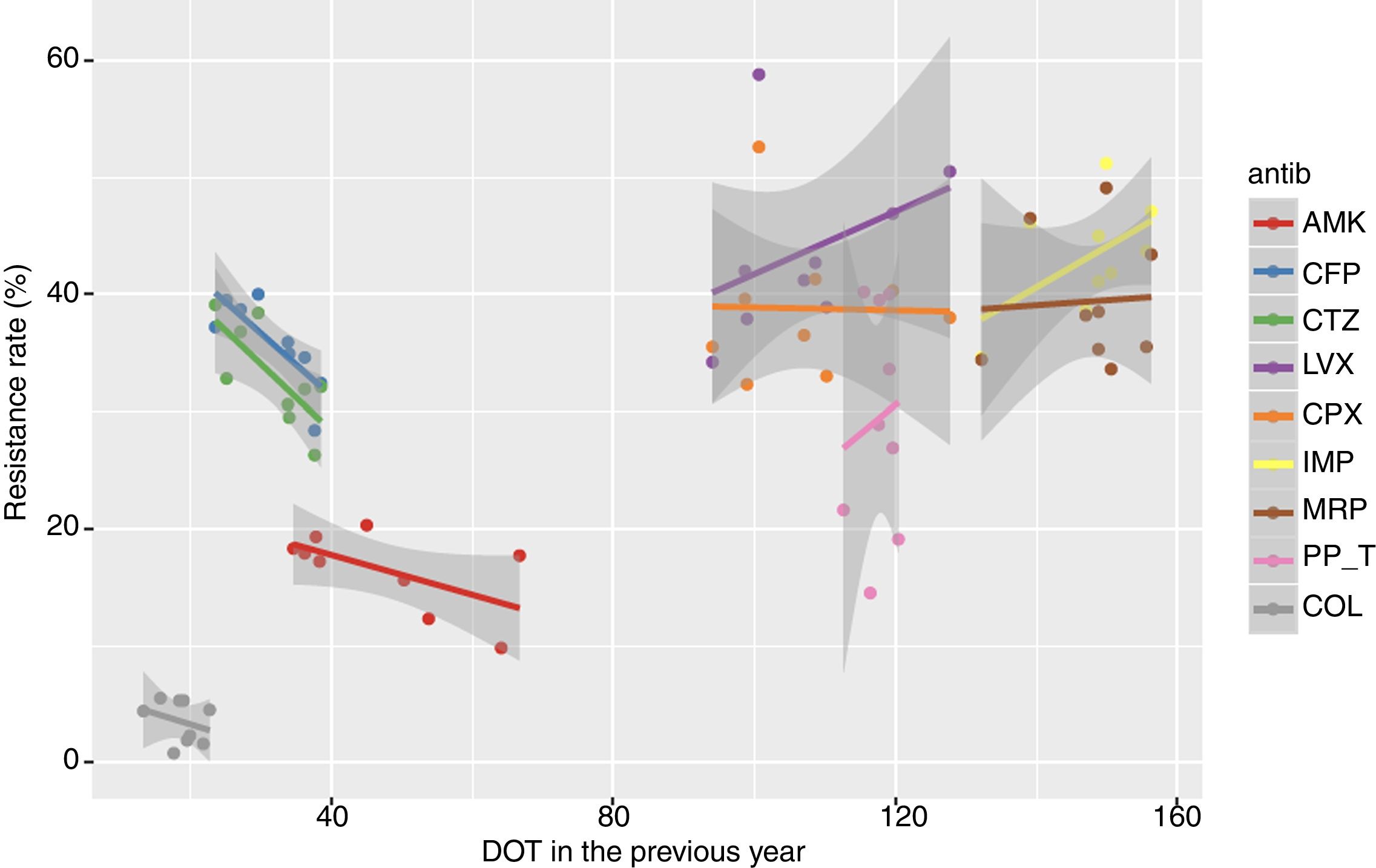
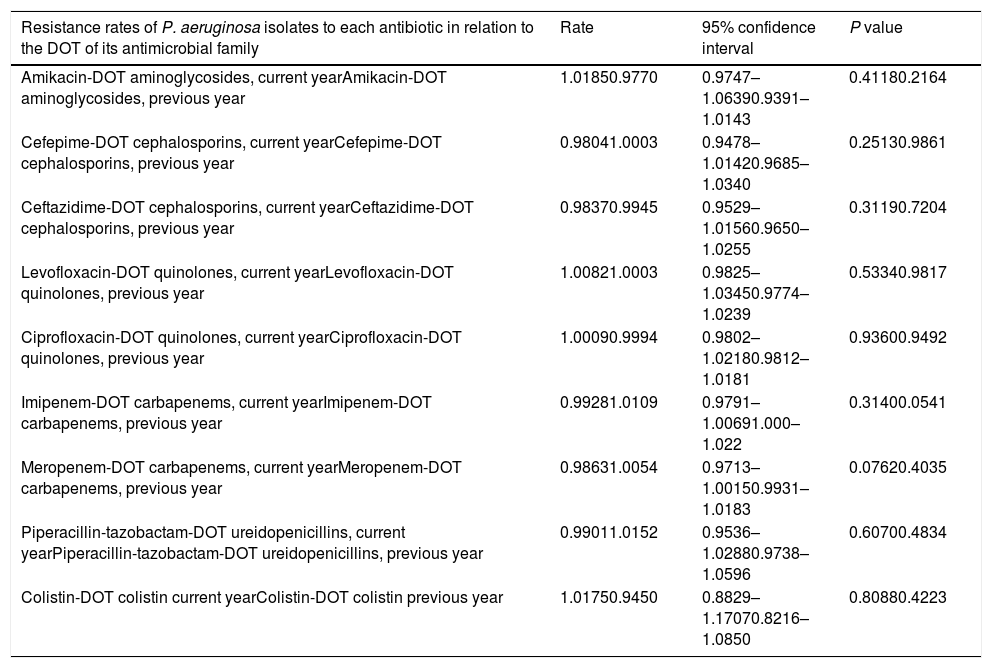
The adjusted Poisson regression model of the relationship between the resistance rates of P. aeruginosa isolates of each antibiotic and the DOT of its antimicrobial family is shown in Table 3. An association between annual resistance rates of P. aeruginosa isolates in ICU-acquired IRDI and consumption antipseudomonal antimicrobial families in the current and previous years was not found. The linear trend of the relationship between resistance rate and DOT for individual antipseudomonal agents and for each antimicrobial is shown in Fig. 3. There was a non-significant inverse association of resistance to ceftazidime and cefepime in the DOT (the higher the consumption, the less the resistance) both of the same year and the previous year, whereas resistance to imipenem and levofloxacin showed a non-significant association with DOT of the same year and the previous year (the higher the consumption, the greater the resistance).
Relationship between evolution of annual resistance rates of P. aeruginosa isolates in ICU-acquired IDRI and consumption of antipseudomonal antimicrobials family in the current and previous years, expressed as DOT (days of treatment/days of ICU stay×1000).
| Resistance rates of P. aeruginosa isolates to each antibiotic in relation to the DOT of its antimicrobial family | Rate | 95% confidence interval | P value |
|---|---|---|---|
| Amikacin-DOT aminoglycosides, current yearAmikacin-DOT aminoglycosides, previous year | 1.01850.9770 | 0.9747–1.06390.9391–1.0143 | 0.41180.2164 |
| Cefepime-DOT cephalosporins, current yearCefepime-DOT cephalosporins, previous year | 0.98041.0003 | 0.9478–1.01420.9685–1.0340 | 0.25130.9861 |
| Ceftazidime-DOT cephalosporins, current yearCeftazidime-DOT cephalosporins, previous year | 0.98370.9945 | 0.9529–1.01560.9650–1.0255 | 0.31190.7204 |
| Levofloxacin-DOT quinolones, current yearLevofloxacin-DOT quinolones, previous year | 1.00821.0003 | 0.9825–1.03450.9774–1.0239 | 0.53340.9817 |
| Ciprofloxacin-DOT quinolones, current yearCiprofloxacin-DOT quinolones, previous year | 1.00090.9994 | 0.9802–1.02180.9812–1.0181 | 0.93600.9492 |
| Imipenem-DOT carbapenems, current yearImipenem-DOT carbapenems, previous year | 0.99281.0109 | 0.9791–1.00691.000–1.022 | 0.31400.0541 |
| Meropenem-DOT carbapenems, current yearMeropenem-DOT carbapenems, previous year | 0.98631.0054 | 0.9713–1.00150.9931–1.0183 | 0.07620.4035 |
| Piperacillin-tazobactam-DOT ureidopenicillins, current yearPiperacillin-tazobactam-DOT ureidopenicillins, previous year | 0.99011.0152 | 0.9536–1.02880.9738–1.0596 | 0.60700.4834 |
| Colistin-DOT colistin current yearColistin-DOT colistin previous year | 1.01750.9450 | 0.8829–1.17070.8216–1.0850 | 0.80880.4223 |
Relationship between resistance rate and DOT of each antipseudomonal antibiotic in the current and the previous year for the period 2008–2016, showing the linear trend for each antibiotic. (AMK: amikacin; CFP: cefepime; CTZ: ceftazidime; LVX: levofloxacin; CPX: ciprofloxacin; IMP: imipenem; MRP: meropenem; PP_T: piperacillin-tazobactam; COL: colistin).
This study failed to demonstrate a statistically significant relationship between the overall consumption of antipseudomonal antimicrobials in ICU populations, expressed as DOT and resistance of P. aeruginosa to individual antimicrobial agent in patients with IDRI related to invasive devices admitted to a large number of Spanish ICUs during a 10-year period. Such negative finding is clinically relevant and in contrast to different studies published in the literature.1,25–34
In individual patients, it has been shown that previous exposure to imipenem or fluoroquinolones influences upon the selection of resistant P. aeruginosa strains to these agents or the development of multiresistant isolates.25–31 In a national surveillance study of antimicrobial consumption in 203 hospitals from Japan, there was a significant association between consumption of imipenem, meropenem, ciprofloxacin, or amikacin and P. aeruginosa resistance to these antimicrobials.32 Data on antimicrobial usage in Korea from 2002 to 2013 revealed that decreasing consumption of amikacin correlated strongly with a decrease of gentamicin-resistant rates of P. aeruginosa.33 Also, in a tertiary-care hospital in Greenville (North Carolina, USA), introduction of ertapenem and reduction of the use of fluoroquinolones was associated with a significant decrease of P. aeruginosa resistant to group 2 carbapenems (imipenem, meropenem, and doripenem).34 In a recent 2-year prospective study conducted in an ICU of a Romanian university hospital,35 the incidence of carbapenem-resistant and multiresistant P. aeruginosa increased significantly, mirroring the increase in consumption of β-lactam agents with β-lactamase inhibitors (piperacillin-tazobactam) and carbapenems (meropenem). However, cross-correlation coefficients and fitted regression models showed that resistance to antimicrobials during a given quarter depends not only on the consumption of antimicrobials during that quarter, but also on consumption during the previous one combined with the incidence of resistant circulating strains.35 By contrast, Fihman et al.36 did not find a correlation between VAP episodes due to P. aeruginosa and antimicrobial consumption in the ICU. Our results of P. aeruginosa strains isolated in ICU-acquired infections related to the use of invasive devices and consumption of antipseudomonal agents during the ICU stay are consistent with these data.
To reduce the presence of multiresistant pathogens, national and international organisms have launched campaigns to optimize the use of antimicrobials in all areas (livestock, agriculture, health sciences),37–39 with emphasis on avoiding the use of antimicrobials in clinical situations where they are not indicated and in reduction of days of treatment.40 In our country, the “Resistant Zero” project developed by the SEMICYUC with the technical support of the Spanish Ministry of Health has been effective to improve antimicrobial use in the ICU setting.41 Recommendations included designation of at least one intensivist as responsible for the use of antimicrobials in each ICU, prioritizing early antimicrobial discontinuation, and limiting empirical coverage with broad-spectrum agents (piperacillin-tazobactam and carbapenems) to severe infections with systemic response (septic shock or severe sepsis).41 All these measures have contributed to reduce ICU consumption of most antimicrobial families with antipseudomonal activity, such as cephalosporins, aminoglycosides, and quinolones except for carbapenems and ureidopenicillins. However, the linear reduction of the use of these antipseudomonal antimicrobials has not been associated with a reduction of the rate of resistance of antipseudomonal antimicrobials in P. aeruginosa strains isolated from ICU-acquired infections in the same year or in the previous year. In a previous study of our group based on data of the ENVIN-HELICS registry over the same 10-year period, a progressive increase of multiresistant isolates, particularly of extensively drug- and pandrug-resistant strains was observed.42 However, in this previous study data on consumption of antipseudomonal antimicrobials was not analyzed.
The presence of MBR strains facilitates the development of a progressively complex endemic flora, which is resistant to common antimicrobial agents and responsible for infections that appear in ICU patients independently of the previously administered antimicrobial treatment. The present findings of a lack of relationship between ICU consumption of antipseudomonal antimicrobials and development of P. aeruginosa resistant strains identified in IDRI related to invasive devices may be explained by different arguments. Firstly, admission to the ICU of patients already colonized with multidrug resistant P. aeruginosa, a status unknown to the medical personnel. These patients can act as a reservoir from which strains are spread through cross-transmission to patients at risk, sometimes without prior mucosal colonization and without prior use of antipseudomonal antimicrobials.43,44 It is recommended to perform active search of MBR pathogens in all patients on admission to the ICU and application of contact precautions in patients at high risk of MBR carriage are some of the recommendations included in the “Resistant Zero” project applied to Spanish ICUs.41 Secondly, there are numerous evidences of MBR reservoirs in the ICU environment (mattress, drains, nebulizers, taps and sinks, portable equipments, floors, walls, etc.) that favor rapid colonization of patients and development of invasive device-associated infections. Active search of MBR pathogens in environmental samples (at least once a week), cleaning protocols with daily cleaning and final cleaning at patient discharge, protocolization of cleaning and disinfection of portable equipments, and application of dry hygienic measures (disposable chlorhexidine towels) in colonized or infected patients are essential to reduce acquisition rates of MBR pathogens.41,45
Limitations of study are its multicenter nature and some design characteristics. Data of identification of P. aeruginosa and susceptibility testing were provided by the Services of Microbiology of the participating hospitals, and a single reference laboratory was not used. On the other hand, a regression to the mean bias cannot be excluded. Also, the transversal nature of the study should be taken into account. A selection bias cannot be excluded because data on consumption of antipseudomonal antimicrobials and resistance rates were based on information of patients admitted during three months (April-June) each year. Likewise, the influence of other factors associated with the selection of MBR strains, such as the concomitant use of other antimicrobials, doses, or sequential therapy45-47 were not assessed since these variables are not registered in the ENVIN database. For this reason, DOT was used instead of daily defined dose (DDD). Finally, different programs aimed to reduce the number ICU-acquired infections due to invasive devices, such “Bacteremia Zero”48 and “Pneumonia Zero”49 as well as “Zero Resistance” program41 could have influenced the decrease of antimicrobial use, although the impact of these interventions cannot be evaluated with the present data.
ConclusionsIn a large number of Spanish ICUs accounting for about 80% of ICUs in the country, a relationship between reduction of consumption of antipseudomonal antimicrobials and resistance rates against most of these agents was not documented. Other factors besides antimicrobial use may account for the increase of resistant P. aeruginosa isolates. Any intervention to reduce MBR P. aeruginosa in the ICU setting should be accompanied by active search of patients carriers of MBR strains on ICU admission together with active search and destroy of potential MBR reservoirs. We suggest that future studies collect detailed information about concomitant patient carriage with P. aeruginosa on admission to intensive care, as well as potential environmental reservoirs.
Consent statementNot applicable given the non-interventional nature of the study because data were collected from the ENVIN-HELICS registry.
Availability of data and materialPlease contact authors for data request.
Authors’ contributionsFAL, conception and design of the study, drafting of the manuscript, data collection, analysis of results, discussion and supervision of the registry; POA, collection and analysis of data, critical review of the manuscript and supervision of the registry; MPM, collection of data and critical review of the manuscript; MC, collection of data and supervision of the registry; XN, collection of data and supervision of the registry; RG, collection of data and supervision of the registry; MPGA, collection of data, interpretation of results and supervision of the registry; ISB, collection of data and supervision of the registry; MMA, statistical analysis and interpretation of results and critical review of the manuscript. All authors have seen and approved the final draft.
FundingNone to be declared.
Conflict of interestsThe authors declare that they have no competing interests.
To all Healthcare professionals, physicians and nurses, who had collaborated in 2007 and 2016 to the ENVIN-HELICS database entering information that has been analyzed in the present study. All of them are coauthors of the study and their names are listed in the annual reports of the ENVIN registry, available at http://hws.vhebron.net/envin-helics/. We are grateful to Sonia Uriona, MD, and Susana Otero, MD, for their contribution in the administration and secretariat of the ENVIN-HELICS registry, and to Marta Pulido, MD, for editing the manuscript and editorial assistance. The fees of medical editing were supported by MSD España. MSD was not involved in the content of the article.